Single-cell analysis of Kaposi’s sarcoma-associated herpesvirus infection in three-dimensional air-liquid interface culture model
- TR Number: 1005
- Authors: Kyle L. Jung, Un Yung Choi, Angela Park, Suan-Sin Foo, Stephanie Kim, Shin-Ae Lee, Jae U. Jung
- Keywords: EpiOral (ORL-606), Kaposi’s sarcoma-associated herpesvirus (KSHV), lytic replication model, virus infection, viral replication and transmission, cell adhesion, cell differentiation, extracellular matrix organization, oxidative stress, apoptosis, PPAR signaling, single cell RNA sequencing (scRNAseq), KSHV infection, Kaposi’s sarcoma (KS), primary effusion lymphoma, multicentric Castleman’s disease, mechanical scoring/ wounding, disruption of epithelial barrier, LANA, ORF65, integrin β1, CD98, RTA, K2, ORF25, KRT5, KRT14, KRT17, S100A2, NOTCH3, LAMB3, S100A4, KRT13, TGM5, S100P, SPRR2A, SPRR2E, keratin 5, keratin 13, keratine 16, S100A7, TGM1, IVL, SFN, EPHA2, GADD45B, CFLAR, RTN4, BAG1, GNB1, RAC1, SDCBP, ATF4, HERPUD1, SOD2, HIF1a, ephrin receptor, NTF2, kallikrein-5 (KLK5), ITGB1, cornulin (CRNN)
- Materials Tested: herpesvirus
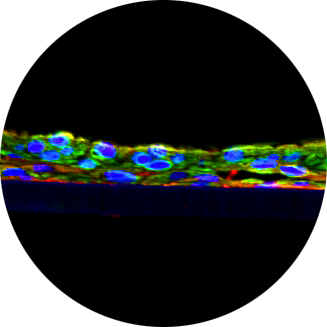
The oral cavity is the major site for transmission of Kaposi’s sarcoma-associated herpesvirus (KSHV), but how KSHV establishes infection and replication in the oral epithelia remains unclear. Here, we report a KSHV spontaneous lytic replication model using fully differentiated, three dimensional (3D) oral epithelial organoids at an air-liquid interface (ALI). This model revealed that KSHV infected the oral epithelia when the basal epithelial cells were exposed by damage. Unlike two-dimensional (2D) cell culture, 3D oral epithelial organoid ALI culture allowed high levels of spontaneous KSHV lytic replication, where lytically replicating cells were enriched at the superficial layer of epithelial organoid. Single cell RNA sequencing (scRNAseq) showed that KSHV infection induced drastic changes of host gene expression in infected as well as uninfected cells at the different epithelial layers, resulting in altered keratinocyte differentiation and cell death. Moreover, we identified a unique population of infected cells containing lytic gene expression at the KSHV K2-K5 gene locus and distinct host gene expression compared to latent or lytic infected cells. This study demonstrates an in vitro 3D epithelial organoid ALI culture model that recapitulates KSHV infection in the oral cavity, where KSHV undergoes the epithelial differentiation-dependent spontaneous lytic replication with a unique cell population carrying distinct viral gene expression.