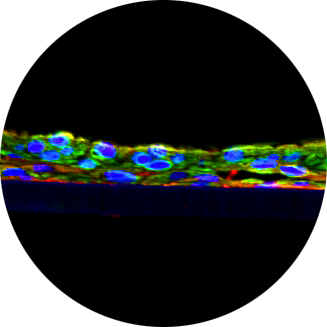
Phenotypic Responses of Differentiated Asthmatic Human Airway Epithelial Cultures to Rhinovirus
- TR Number: 760
- Authors: Jianwu Bai1*, Steven L. Smock1, George R. Jackson Jr.2 Kenzie D. MacIsaac1, Yongsheng Huang1, Courtney Mankus2, Jonathan Oldach2, Brian Roberts1¤a, Yu-Lu Ma1, Joel A. Klappenbach1, Michael A. Crackower1¤b, Stephen E. Alves1, Patrick J. Hayden2* 1 Merck Research Laboratories, Boston, Massachusetts, United States of America, 2 MatTek Corporation, Ashland, Massachusetts, United States of America ¤a Current address: HudsonAlpha Institute for Biotechnology, Huntsville, Alabama, United States of America ¤b Current address: Biogen Idec, Cambridge, Massachusetts, United States of America
- Keywords: Airway epithelial monolayers, Asthma, bioinformatics analysis, differential gene expression, EpiAirway (AIR-100), EpiAirway (AIR-100-D2), human rhinovirus, Muc5AC, RNA- seq, viral exacerbation
- Materials Tested: HRV16, poly(I:C), rhinovirus
Objectives: Human airway epithelial cells are the principal target of human rhinovirus (HRV), a common cold pathogen that triggers the majority of asthma exacerbations. The objectives of this study were 1) to evaluate an in vitro air liquid interface cultured human airway epithelial cell model for HRV infection, and 2) to identify gene expression patterns associated with asthma intrinsically and/or after HRV infection using this model. Methods: Air-liquid interface (ALI) human airway epithelial cell cultures were prepared from 6 asthmatic and 6 non-asthmatic donors. The effects of rhinovirus RV-A16 on ALI cultures were compared. Genome-wide gene expression changes in ALI cultures following HRV infection at 24 hours post exposure were further analyzed using RNA-seq technology. Cellular gene expression and cytokine/chemokine secretion were further evaluated by qPCR and a Luminex-based protein assay, respectively. Main Results: ALI cultures were readily infected by HRV. RNA-seq analysis of HRV infected ALI cultures identified sets of genes associated with asthma specific viral responses. These genes are related to inflammatory pathways, epithelial structure and remodeling and cilium assembly and function, including those described previously (e.g. CCL5, CXCL10 and CX3CL1, MUC5AC, CDHR3), and novel ones that were identified for the first time in this study e.g.CCRL1).