Microbially competent 3D skin: a test system that reveals insight into host–microbe interactions and their potential toxicological impact
- TR Number: 917
- Authors: Lisa Lemoine, Ralf Dieckmann, Sascha Al Dahouk, Szilvia Vincze, Andreas Luch, Tewes Tralau
- Keywords: EpiDermFT (EFT-400), skin microbiome, Micrococcus luteus, Pseudomonas oleovorans, IL-1a, IL-1b, IL-6, antimicrobial peptides, TLR2, HGF, EGF, IGFBP-1, IGFBP-4, IGFBP-5, IGFBP-6, PDGFA, PDGFD, CXCL2, CXCL3, CXCL8, CXCL12, FGF5, TGFB1, VEGFA, VEGFC, FGF4, FGF2, CYP1A1, CYP1B1, CYP2D6, CYP4B1, CYP2J2, CYP4F12,, CYP26B1, CYP2C18, CYP24A1, CYP27A1, CYP19A1, CYP4V2, CYP3A5, CYP3A7, CYP7B1, CYP26A1, CYP2A6, CYP2F1, Defensin b4a, IL-1ra, C5/C5a, G-CSF, GM-CSF, CD54, IL-8, GRO-a, MCP-1, MIF, Serpin E1, IL-37, IL-36B, IL-36G, IL-10, IL-19, IL-23A, IL-24, DEFB4B, DEFB4A, DEFB103A, DEFB103B, CSF2, CSF3, IL1RN, CCL3
- Materials Tested: M. luteus, P. oleovorans
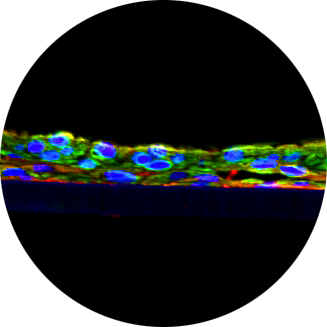
The skin`s microbiome is predominantly commensalic, harboring a metabolic potential far exceeding that of its host. While there is clear evidence that bacteria-dependent metabolism of pollutants modulates the toxicity for the host there is still a lack of models for investigating causality of microbiome-associated pathophysiology or toxicity. We now report on a biologically characterized microbial–skin tissue co-culture that allows studying microbe–host interactions for extended periods of time in situ. The system is based on a commercially available 3D skin model. In a proof-of-concept, this model was colonized with single and mixed cultures of two selected skin commensals. Two different methods were used to quantify the bacteria on the surface of the skin models. While Micrococcus luteus established a stable microbial–skin tissue co-culture, Pseudomonas oleovorans maintained slow continuous growth over the 8-day cultivation period. A detailed skin transcriptome analysis showed bacterial colonization leading to up to 3318 significant changes. Additionally, FACS, ELISA and Western blot analyses were carried out to analyze secretion of cytokines and growth factors. Changes found in colonized skin varied depending on the bacterial species used and comprised immunomodulatory functions, such as secretion of IL-1α/β, Il-6, antimicrobial peptides and increased gene transcription of IL-10 and TLR2. The colonization also influenced the secretion of growth factors such as VFGFA and FGF2. Notably, many of these changes have already previously been associated with the presence of skin commensals. Concomitantly, the model gained first insights on the microbiome’s influence on skin xenobiotic metabolism (i.e., CYP1A1, CYP1B1 and CYP2D6) and olfactory receptor expression. The system provides urgently needed experimental access for assessing the toxicological impact of microbiome-associated xenobiotic metabolism in situ.